 Home > Achievement > Research Project Summary Report > 2017 year > The application of Single Nucleotide Polymorphism (SNP) markers in identification of resistance in tomato and hybrid seed purity in watermelon. Home > Achievement > Research Project Summary Report > 2017 year > The application of Single Nucleotide Polymorphism (SNP) markers in identification of resistance in tomato and hybrid seed purity in watermelon. |
The application of Single Nucleotide Polymorphism (SNP) markers in identification of resistance in tomato and hybrid seed purity in watermelon.
In this study, the tomato resistance-related gene sequences of Sw-5 ,Tm-1,Tm-22 ,Ph-2 and Ph-3 genes were used to design SNP primers. The resistance and susceptible tomato materials were screened by using PCR and real-time PCR. Then the screening results showed that these tomato materials could be divided into three resistance genotypes in Sw-5 ,Tm-1 ,Tm-22, Ph-2 and Ph-3 genes. In addition, we had done the molecular detection and field traits with two-hybrid tomato F1 materials from KNOWYOU SEED company. We selected 20 single plants with resistance genes and good characteristics based on the analyzed results. Seven single plants of them were licensed to three companies. Analyzed 6,784,860 SNPs from public genome data of watermelon, 2,156 SNPs whose minor allele frequency (MAF) were more than 0.47 were selected and designed to 857 allele specific markers among which 140 AS markers were synthesized and tested. The results showed that 63 AS markers are polymorphic among the tested hybrid varieties as hybrid seed purity test markers. It was expected that these markers could be widely used in other watermelon varieties.
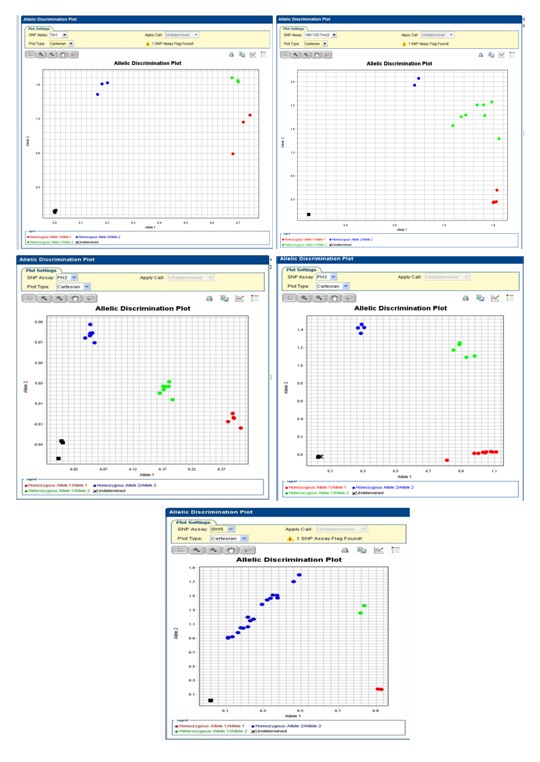
▲Fig. 1 The SNP molecular markers linked to resistant
genes of tomato mosaic virus, Phytophthora infestans
and tomato spotted wilt virus developed by TSIP.
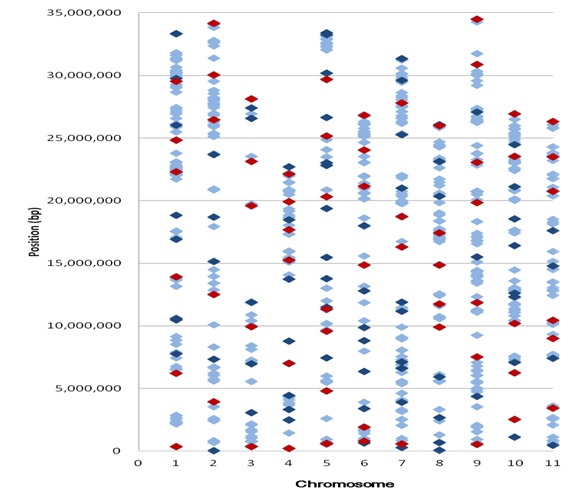
▲Fig. 2 Distribution of SNPs designed as the allele-specific (AS)
markers on the genome of watermelon. The light blue spots were
857 SNPs initially selected; the dark blue and red spots were the
140 SNP (AS) markers tested in the electrophoresis analysis; and
the red spots were the 63 polymorphic candidate SNPs developed
in this study.
