2025 Research Project List
首頁 > 英文版網站 > Achievement > 2025 Research Project List > Development and Application of Molecular Markers Associated with Anti-fusarium Wilt on Luffa and Drought-tolerance and Other Important Traits on Maize
Development and Application of Molecular Markers Associated with Anti-fusarium Wilt on Luffa and Drought-tolerance and Other Important Traits on Maize
This project consists of two research subprojects:
- Utilization and development of molecular markers associated with drought tolerance and other important traits in sweet corn: The drought-tolerant hybrid population of edible corn continues to advance through self-crossing generations, aiming to obtain Recombinant Inbred Lines (RILs) for deeper genetic analysis and to be breeding materials. In addition, a hybrid combination (ZM-00003 × ZM-00004) of potential drought-tolerant varieties was utilized as the base material for advancing selfing generations and conducting related investigative analyses this year. According to the SNP polymorphism analysis of drought-associated genes (dhn1 and rsp41), a total of 15 individual plants in the S1 generation were identified as possessing potential drought-tolerant characteristics. Furthermore, the analysis of SNP loci in the sh2 gene indicated a high correlation between the molecular marker polymorphism and the ear appearance (shrunken/full) of the lines. However, the observed correlation was entirely opposite to the findings of previous studies. This unexpected finding necessitates futher observation to resolve the discrepancy.
- Development of molecular markers for fusarium wilt resistance in Luffa: a candidate SNP molecular marker for fusarium wilt resistance located on chromosome 6 was successfully designed this year. However, inoculation trials indicated that the ambient temperature significantly affects the incidence of fusarium wilt, resulting in a low prediction accuracy for this candidate marker. To overcome the influence of environmental variation on QTL detection power, subsequent plans involve re-inoculating the remaining F2:3 line seeds and utilizing the F2:3 disease resistance ratio as the phenotype to enhance the power of gene mapping.
![Figure 1. PCR electrophoresis results using SNP allele-specific primers [T/C]. (a) Genotyping results of the resistant and susceptible parental lines and the F1 generation; (b) Genotyping results of 12 F2:3 plants (line No. 006).](/upload/tss/images/web_structure/8273/29-01(1).jpg) ▲Figure 1. PCR electrophoresis results using SNP allele-specific primers [T/C]. (a) Genotyping results of the resistant and susceptible parental lines and the F1 generation; (b) Genotyping results of 12 F2:3 plants (line No. 006). |
.jpg) ▲Figure 2. The second inoculation test (October 17, 2025) of F2:3 lines derived from crosses of resistant and susceptible luffa lines for Fusarium wilt. The right panel shows successful disease development in the susceptible control ‘Yinguang’, while the left panel shows segregation in the inoculation response of one F2:3 line. |
|
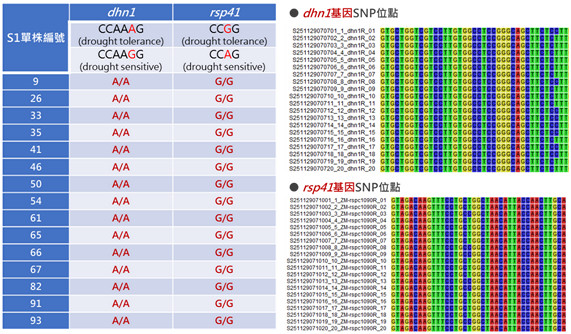 ▲Figure 3. Molecular marker (dhn1, rsp41) analysis of the S1 generation derived from the maize hybrid combination (ZM-00003 × ZM-00004). |
 ▲Figure 4. Association between the sh2 gene SNP locus in individual S1 plants and the appearance of their selfed ears (S2). |
|
© www.tss.gov.tw
https://www.tss.gov.tw/ws.php?id=8273
2026/04/08
 農業部種苗改良繁殖場
農業部種苗改良繁殖場