2025 Research Project List
This project aims to comprehensively assess the current status of grapevine viruses in Taiwan, enhance diagnostic efficiency, and strengthen the production system for virus-free planting materials. During 2024-2025, a total of 164 grapevine samples were collected across four counties and seven major production areas. Using small RNA next-generation sequencing (sRNA-NGS) combined with molecular diagnostics, we successfully characterized the viral communities present in the field and verified the presence of multiple viruses within infected plants. This approach revealed several previously overlooked latent viruses. A total of seven grapevine viruses were detected in Taiwan vineyards (GVA, GLRaV-1, GLRaV-3, GFkV, GFabV, GGVA, and GVE). Among these, Grapevine geminivirus A (GGVA) was the most prevalent and severe, with infection rates reaching 70-100% across the seven major production regions. The Xihu area showed the highest incidence (100%), followed by Neiwan (96%), Zhuolan (90%), and Fengqiu (86%), all exhibiting high levels of infection. Notably, GFkV and GLRaV-3 also showed elevated incidence rates in certain regions. Mixed viral infections were widespread, with samples from Xinshe, Dacun, and Neiwan frequently harboring five viruses simultaneously. A previous survey by Yang et al. (2000) reported post-planting infections of GLRaV-1, GFLV, and GVA in grapevine plantlets, but the viral community has changed significantly over the past two decades. Recent investigations identified three additional viruses (GFabV, GVE, and GGVA) now present in Taiwan vineyards, indicating ongoing shifts and evolution within viral populations. This year, we continued to extract nucleic acids from tissue cultured plantlets of six cultivars maintained at our station, pooled the samples, and performed small RNA NGS sequencing. The results suggested that two cultivars collected externally were virus-infected. Follow-up confirmation using specific primers showed that rootstock 420A was infected with GFabV, while the Muscat cultivar carried GGVA, with sequence identity confirmed through alignment. No target viruses were detected in cultivars 5C, 8B, Sakurai, or Isseki. Overall, this project has clarified the threat level, distribution patterns, and mixed infection status of grapevine viruses in Taiwan. These findings will guide future improvements in viral diagnostics and the production system for virus-free planting materials, laying an essential foundation for disease management and phytosanitary security in Taiwan’s grapevine industry.
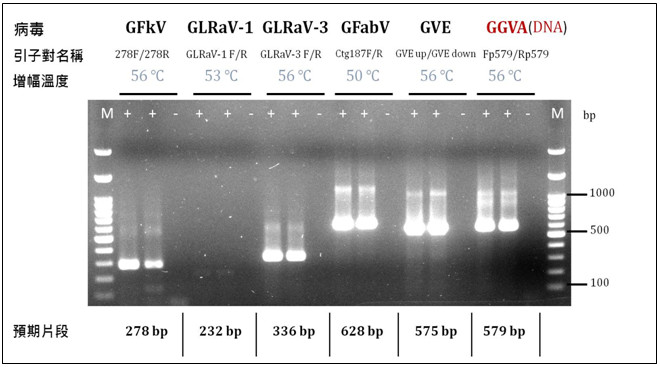 ▲Figure 1. Development of molecular detection methods for six grapevine viruses. |
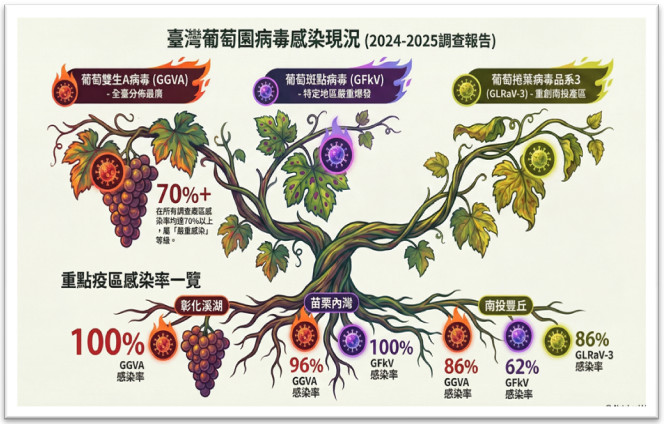 ▲Figure 2. Current status of grapevine virus infections in the field during 2024-2025. |


